 We’re back to genetics, and Jeffrey Tomkins thinks he’s disproved evolution once again in Genetic Recombination Study Defies Human-Chimp Evolution. He says:
We’re back to genetics, and Jeffrey Tomkins thinks he’s disproved evolution once again in Genetic Recombination Study Defies Human-Chimp Evolution. He says:
Results from a recent study in human and chimpanzee genetics have shipwrecked yet another Darwinian hypothesis.
Some parts of the chimp and human genomes are more similar to each other than others. According to Tomkins, the process of genetic recombination is supposed to have produced these differences. He points to a new paper – Recombination Rates and Genomic Shuffling in Human and Chimpanzee—A New Twist in the Chromosomal Speciation Theory – that shows, among other things, that recombination rates are actually lower in places where there are more differences. Or, in his own words:
These results are the exact opposite of what evolutionists expected. According to evolutionary reasoning, the chromosomal areas between humans and chimps that were the most different should have had high levels of genetic recombination that would help explain why they were so different. But these chromosomal areas that were the most different between humans and chimpanzees had the lowest levels!
More recombination equals more evolutionary differences right? Apparently not!
At first glance, this is a convincing argument. But there is a loose end here, from which the whole story unravels: why is there a correlation here at all? Creationists acknowledge that some animals are more similar than others, but try to explain this away by the need for common function producing common design. If recombination produces change, why are the parts that are supposed to be most similar changing the most? Even if it doesn’t, that still doesn’t explain why there should be any difference at all. The creationist prediction here should really be that all parts of the genome express recombination equally, but certainly not that the “common design” should be affected most whatever its consequences.
To see how this loose end ruins Tomkins argument requires first taking a look at what recombination actually is. Tomkins describes it like so:
When sperm and egg cells are formed in humans and various animals, the process of meiosis generates genetic variation. For example, since humans have two sets of chromosomes, when similar ones (i.e., sister chromatids)–one each from your mother and father—pair up together in the cell, they undergo a controlled exchange of DNA segments (maintaining the same linear order of segments). This is one reason why the offspring of two parents are always genetically unique, except for identical twins where the fertilized egg cell splits into two identical embryos. This process of exchanging DNA segments across sister chromatids is called genetic or homologous recombination and does not occur randomly across the genome, but most often occurs in areas called “hotspots.”
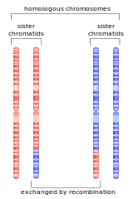
Technically speaking, homologous recombination is just one type of genetic recombination. The concept is summed up in the large scale by the picture above and to the right, but at the small scale it is important to note that recombination is the result of fixing breaks in the chromosome – sometimes the break is fixed simply by sticking the broken bits back together again, while other times the broken part gets stuck to the wrong chromosome in the pair with the net affect of swapping huge sections of each chromosome with each other. The process does produce new variation in a sense: if we imagine that the red chromosome held the alleles “AB,” and the blue “ab,” recombination can potentially add “Ab” and “aB” to the mix, though it generally wont produce a “C.” Other forms of large-scale chromosomal change beyond simple homologous recombination can give you outcomes like “AAB” and “ba,” but we’ll get to the affects of that later.
The hotspots that Tomkins talks about are the consequence of the fact that the breaks that lead to recombination are significantly more likely to occur in certain short regions of a few thousand base pairs than in the rest of the genome – see here for a graph showing what this looks like. Around a year ago Tomkins was claiming that this fact alone “negated” evolution, but this is not the case. In addition, while recombination of some form is a constant among all life (even viruses, apparently) the famous fruit fly Drosophila melanogaster is one of an number of species that do not have hotspots at all.
If recombination acts to blend “AB” and “ab” into “Ab” and “aB,” and is a mechanism for fixing breakages, then it’s not a very good way of driving species apart now is it? Quite the contrary, in fact: as you might remember from discussions of Mendelian inheritance in biology class, “blending” is if anything a problem for evolution making progress. But if you don’t have so much recombination chromosomes can mutate far from the average state without being blended back. Lower levels of recombination, therefore, can be expected to lead to greater, not lesser, differences between chromosomes and species. Case closed then, right? Not quite.
According to Tomkins, the research in the paper he is discussing found:
The researchers found that genetic recombination levels were much higher in regions of the genome between humans and chimps where sequence identity was higher. In the regions of much lower DNA similarity, which occur as differences in gene order, gene content, and other major DNA sequence differences—the recombination rates were much lower.
This is not quite true: rather than comparing regions based on all of “differences in gene order, gene content, and other major DNA sequence differences” the study specifically looked at chromosomes based on whether they had been rearranged – i.e. changes in gene order and direction, but not changes within the genes themselves. They found that when chromosomes had been rearranged they exhibited much lower levels of recombination. This supports the hypothesis that they were testing, which is that rearrangements within chromosomes suppress recombination and thus help drive speciation.
The take-home summary is that Tomkins has the cause and effect in this story all wrong. He thinks that recombination should cause differences, and that the fact that the opposite correlation is observed “has falsified a prominent evolutionary hypothesis.” But instead the result supports the hypothesis that it is chromosomal rearrangements that cause a reduction in recombination. That’s a long way from being a blow to evolution, but I still want to see how it can even be made to fit into the creationist view.
There is another claim that Tomkins makes in his article that doesn’t seem to be directly related:
Interestingly, the authors also searched the DNA sequences between humans and chimpanzees for sections that were “flipped” in their orientation, called inversions. Large inversions, once they occur in a species and if they are tolerated, will stop recombination. However, the researchers found that inverted sequences accounted for very few differences in the regions they examined.
“[I]nverted sequences accounted for very few differences” between what? It isn’t clear what Tomkins means here – does inversion not change much about the sequence, or the rate of recombination? From my reading, neither statement is actually true (so long as you count the same sequence going in a completely different direction as significantly “different”), but maybe he means something completely different?

Interesting how the comment at the end of the Abstract that the scientists’ observations provide “new evidences for the contribution of inversions in suppressing recombination in mammals” has transformed into Tomkins’ title which claims this study “defies Human-Chimp Evolution” (a highly misleading comment anyway – if it is intended to imply that human species evolved FROM chimps rather than that both diverged from a common ancestor).
Just so you have it, should someone came searching …
PZ Myers just did a refutation of Tompkins (complete with pretty diagrams) here:
http://freethoughtblogs.com/pharyngula/2013/06/18/do-the-creationist-shuffle-and-twist/
Just noticed John Pieret’s very recent comment, ahead of posting mine.
Only somewhat hurriedly skimmed (as previously) but it seems that an atheist and a YEC have reached contrasting conclusions about claims in a recent pro-Christian article that quotes Jeffrey Tomkins and another YEC:
http://freethoughtblogs.com/pharyngula/2013/06/18/do-the-creationist-shuffle-and-twist/#more-11625
http://answersforhope.com/evolution-and-monkey-business-a-recent-study-poses-new-problems-for-evolution/
This is the article:
http://christiannews.net/2013/06/17/groundbreaking-genetic-discoveries-challenge-ape-to-human-evolutionary-theory/
This is the Abstract of the paper in question:
Click to access mss272.pdf
I haven’t studied this closely, but what appears to have been brought into question (according to ‘Eye’ and to Myers) is what creationists THINK evolutionists believed before this paper was published.